(+)-Isocorynoline
Modify Date: 2025-08-23 12:27:01
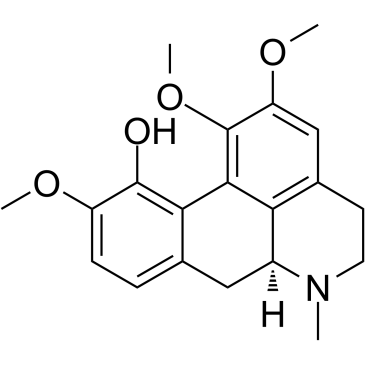
(+)-Isocorynoline structure
|
Common Name | (+)-Isocorynoline | ||
|---|---|---|---|---|
| CAS Number | 475-67-2 | Molecular Weight | 341.401 | |
| Density | 1.2±0.1 g/cm3 | Boiling Point | 506.1±50.0 °C at 760 mmHg | |
| Molecular Formula | C20H23NO4 | Melting Point | 216-220ºC(lit.) | |
| MSDS | N/A | Flash Point | 259.9±30.1 °C | |
Use of (+)-IsocorynolineIsocorydine is isolated from Dicranostigma leptopodum (Maxim.) Fedde (DLF). Isocorydine combines with Doxorubicin (DOX) has a promising potential to eradicate hepatocellular carcinoma (HCC)[1]. |
| Name | isocorydine hydrochloride |
|---|---|
| Synonym | More Synonyms |
| Description | Isocorydine is isolated from Dicranostigma leptopodum (Maxim.) Fedde (DLF). Isocorydine combines with Doxorubicin (DOX) has a promising potential to eradicate hepatocellular carcinoma (HCC)[1]. |
|---|---|
| Related Catalog | |
| In Vitro | Isocorydine (0-400 ug/ml; 48 hours) show a significant decrease in the IC50 for ICD and DOX, the CI values are 0.605, 0.644, 0.804, and 0.707 respectively for Huh-7, Hep-G2, SNU-449 and SNU-387[1]. Isocorydine (0-400 ug/ml; 48 hours) abrogates DOX-induced upregulation of mesenchymal markers and the downregulation of epithelial markers in human HCC cell lines[1]. Cell Viability Assay[1] Cell Line: Huh-7, Hep-G2 , SNU-387, SNU-449 cells Concentration: 0-400 ug/ml Incubation Time: 24 hours Result: Had a higher cytotoxicity in HCC cells in comparison to ICD or DOX alone. Western Blot Analysis[1] Cell Line: Huh-7, Hep-G2 , SNU-387, SNU-449 cells Concentration: Incubation Time: 24 hours Result: Downregulated protein levels of Claundin-1 and E-cadherin. |
| In Vivo | Isocorydine (intraperitoneal injection; 0.4 mg/kg; every 2 days for 2 weeks) retards the tumor growth, but the combined treatment of Doxorubicin (DOX) or ICD significantly inhibits tumor growth[1]. Animal Model: Female nude mice[1] Dosage: 0.4 mg/ml Administration: Injected intraperitoneally every 2 days for 2 weeks Result: Combined treatment of isocorydine and DOX showed a promising potential to eradicate HCC. |
| References |
| Density | 1.2±0.1 g/cm3 |
|---|---|
| Boiling Point | 506.1±50.0 °C at 760 mmHg |
| Melting Point | 216-220ºC(lit.) |
| Molecular Formula | C20H23NO4 |
| Molecular Weight | 341.401 |
| Flash Point | 259.9±30.1 °C |
| Exact Mass | 341.162720 |
| PSA | 51.16000 |
| LogP | 3.09 |
| Vapour Pressure | 0.0±1.4 mmHg at 25°C |
| Index of Refraction | 1.604 |
| InChIKey | QELDJEKNFOQJOY-ZDUSSCGKSA-N |
| SMILES | COc1ccc2c(c1O)-c1c(OC)c(OC)cc3c1C(C2)N(C)CC3 |
| Storage condition | -20℃ |
CHEMICAL IDENTIFICATION
HEALTH HAZARD DATAACUTE TOXICITY DATA
|
| WGK Germany | 3 |
|---|---|
| RTECS | CE1057950 |
|
Name: Luminescence-based cell-based primary high throughput screening assay to identify ago...
Source: The Scripps Research Institute Molecular Screening Center
Target: mu-type opioid receptor isoform MOR-1 [Homo sapiens]
External Id: OPRM1-OPRD1_AG_LUMI_1536_1X%ACT PRUN
|
|
Name: QFRET-based biochemical primary high throughput screening assay to identify exosite i...
Source: The Scripps Research Institute Molecular Screening Center
Target: disintegrin and metalloproteinase domain-containing protein 17 preproprotein [Homo sapiens]
External Id: ADAM17_INH_QFRET_1536_1X%INH PRUN
|
|
Name: HepG2 viability counterscreen (readout 2: ATP content) against the NCATS DSHEA and TC...
Source: NCGC
Target: N/A
External Id: DSHEA-v1-HepG2-viability-CTG-ATP
|
|
Name: ERK5 transcriptional activity HTS
Source: 24565
Target: N/A
External Id: ERK5 transcriptional activity-HTS
|
|
Name: Fluorescence-based cell-based primary high throughput screening assay to identify ago...
Source: The Scripps Research Institute Molecular Screening Center
Target: muscarinic acetylcholine receptor M1 [Homo sapiens]
External Id: CHRM1_AG_FLUO8_1536_1X%ACT PRUN
|
|
Name: uHTS identification of small molecule activators of the adaptive arm of the Unfolded ...
Source: Burnham Center for Chemical Genomics
Target: N/A
External Id: BCCG-A405-UPR-XBP1-PrimaryAgonist-Assay
|
|
Name: Fluorescence polarization to screen for inhibitor that competite the binding of FadD2...
Source: Broad Institute
Target: FATTY-ACID-CoA LIGASE FADD28 (FATTY-ACID-CoA SYNTHETASE)
External Id: 2147-01_Inhibitor_SinglePoint_HTS_Activity
|
|
Name: Bursicon-induced LGR2 mediated cAMP production in LGR-2/CRE6x-Luciferase co-transfect...
Source: Broad Institute
Target: N/A
External Id: Bursicon-induced LGR2 mediated cAMP production in LGR-2/CRE6x-Luciferase co-transfected HEK293 cells Inhibition - 7011-01_Antagonist_SinglePoint_HTS_Activity
|
|
Name: Primary Screen Inhibitors of CD40 Signaling in BL2 Cells Measured in Cell-Based Syste...
Source: Broad Institute
Target: N/A
External Id: 7124-01_Inhibitor_SinglePoint_HTS_Activity
|
|
Name: Fluorescence-based cell-based primary high throughput screening assay to identify pos...
Source: The Scripps Research Institute Molecular Screening Center
Target: muscarinic acetylcholine receptor M1 [Homo sapiens]
External Id: CHRM1_PAM_FLUO8_1536_1X%ACT PRUN
|
Total 153, Current Page 1 of 16
1
2
3
4
5
| artabotrin |
| 1,2,10-Trimethoxy-6aa-aporphin-11-ol |
| d-isocorydine |
| (6aS)-1,2,10-Trimethoxy-6-methyl-5,6,6a,7-tetrahydro-4H-dibenzo[de,g]quinolin-11-ol |
| luteanin |
| isocorydine |
| (S)-5,6,6a,7-Tetrahydro-1,2,10-trimethoxy-6-methyl-4H-dibenzo[de,g]quinolin-11-ol |
| iso-Corydine |
| LUTEANINE |
 CAS#:50-00-0
CAS#:50-00-0![4,5,6aα,7-Tetrahydro-1,2,10-trimethoxy-6H-dibenzo[de,g]quinoline-11-ol Structure](https://image.chemsrc.com/caspic/138/475-70-7.png) CAS#:475-70-7
CAS#:475-70-7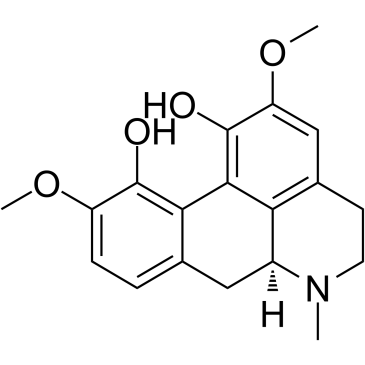 CAS#:517-56-6
CAS#:517-56-6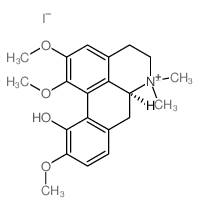 CAS#:17669-18-0
CAS#:17669-18-0 CAS#:13552-72-2
CAS#:13552-72-2
