Epiberberine
Modify Date: 2025-08-25 11:58:01
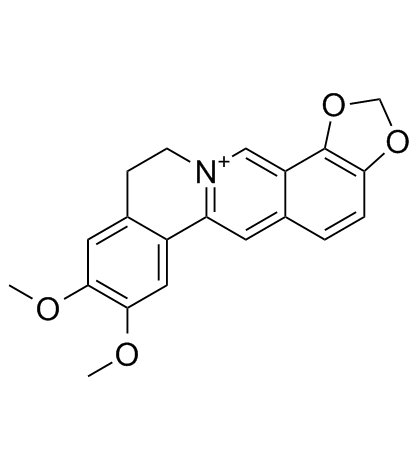
Epiberberine structure
|
Common Name | Epiberberine | ||
|---|---|---|---|---|
| CAS Number | 6873-09-2 | Molecular Weight | 336.36 | |
| Density | N/A | Boiling Point | N/A | |
| Molecular Formula | C20H18NO4+ | Melting Point | 260ºC | |
| MSDS | N/A | Flash Point | N/A | |
Use of EpiberberineEpiberberine is an alkaloid isolated from Coptis chinensis, acts as a potent AChE and BChE inhibitor, and a non-competitive BACE1 inhibitor, with IC50s of 1.07, 6.03 and 8.55 μM, respectively. Epiberberine has antioxidant activity, with peroxynitrite ONOO- scavenging effect (IC50, 16.83 μM), and may protect against Alzheimer disease[1]. Epiberberine inhibits the early stage of differentiation of 3T3-L1 preadipocytes, downregulates the Raf/MEK1/2/ERK1/2 and AMPKα/Akt pathways[2]. Epiberberine has the potential effect in the research of diabetic disease[3]. |
| Name | Epiberberine |
|---|---|
| Synonym | More Synonyms |
| Description | Epiberberine is an alkaloid isolated from Coptis chinensis, acts as a potent AChE and BChE inhibitor, and a non-competitive BACE1 inhibitor, with IC50s of 1.07, 6.03 and 8.55 μM, respectively. Epiberberine has antioxidant activity, with peroxynitrite ONOO- scavenging effect (IC50, 16.83 μM), and may protect against Alzheimer disease[1]. Epiberberine inhibits the early stage of differentiation of 3T3-L1 preadipocytes, downregulates the Raf/MEK1/2/ERK1/2 and AMPKα/Akt pathways[2]. Epiberberine has the potential effect in the research of diabetic disease[3]. |
|---|---|
| Related Catalog | |
| Target |
IC50: 1.07 μM (AChE), 6.03 μM (BChE), 8.55 μM (BACE1)[2] |
| In Vitro | Epiberberine (0, 12.5, 25, or 50 μM) dose-dependently inhibits cellular triglyceride accumulation in 3T3-L1 adipocytes, with an IC50 of 52.8 μM[2]. Epiberberine (12.5-50 μM) suppresses the Raf/MEK1/ERK1/2 and AMPKα/Akt pathways in the early stage of 3T3-L1 adipocyte differentiation[2]. Epiberberine (0.2, 1, 5 μg/mL) inhibits glucose uptake in HepG2 cells in a concentration-dependent manner[3]. |
| In Vivo | Epiberberine (225 mg/kg, p.o. daily for 40 days) reduces body weight, food consumption, water intake, and urinary output of KK-Ay mice[3]. |
| References |
| Melting Point | 260ºC |
|---|---|
| Molecular Formula | C20H18NO4+ |
| Molecular Weight | 336.36 |
| PSA | 40.80000 |
| LogP | -0.99 |
| InChIKey | FPJQGFLUORYYPE-UHFFFAOYSA-N |
| SMILES | COc1cc2c(cc1OC)-c1cc3ccc4c(c3c[n+]1CC2)OCO4 |
| Storage condition | 2-8C |
|
Name: Primary cell-based high-throughput screening assay for identification of compounds th...
Source: Johns Hopkins Ion Channel Center
Target: regulator of G-protein signaling 4 isoform 2 [Homo sapiens]
External Id: JHICC_RGS_Act_HTS
|
|
Name: Luminescence-based cell-based primary high throughput screening assay to identify ago...
Source: The Scripps Research Institute Molecular Screening Center
Target: mu-type opioid receptor isoform MOR-1 [Homo sapiens]
External Id: OPRM1-OPRD1_AG_LUMI_1536_1X%ACT PRUN
|
|
Name: QFRET-based biochemical primary high throughput screening assay to identify exosite i...
Source: The Scripps Research Institute Molecular Screening Center
Target: disintegrin and metalloproteinase domain-containing protein 17 preproprotein [Homo sapiens]
External Id: ADAM17_INH_QFRET_1536_1X%INH PRUN
|
|
Name: Fluorescence-based cell-based primary high throughput screening assay to identify ago...
Source: The Scripps Research Institute Molecular Screening Center
Target: muscarinic acetylcholine receptor M1 [Homo sapiens]
External Id: CHRM1_AG_FLUO8_1536_1X%ACT PRUN
|
|
Name: uHTS identification of small molecule activators of the adaptive arm of the Unfolded ...
Source: Burnham Center for Chemical Genomics
Target: N/A
External Id: BCCG-A405-UPR-XBP1-PrimaryAgonist-Assay
|
|
Name: High throughput fluorescence intensity-based biochemical assay to screen for small mo...
Source: University of Pittsburgh Molecular Library Screening Center
Target: furin (paired basic amino acid cleaving enzyme), isoform CRA_a [Homo sapiens]
External Id: MH080376 Biochemical HTS for Inhibitors of the Proprotein Convertase Furin.
|
|
Name: Fluorescence polarization to screen for inhibitor that competite the binding of FadD2...
Source: Broad Institute
Target: FATTY-ACID-CoA LIGASE FADD28 (FATTY-ACID-CoA SYNTHETASE)
External Id: 2147-01_Inhibitor_SinglePoint_HTS_Activity
|
|
Name: Bursicon-induced LGR2 mediated cAMP production in LGR-2/CRE6x-Luciferase co-transfect...
Source: Broad Institute
Target: N/A
External Id: Bursicon-induced LGR2 mediated cAMP production in LGR-2/CRE6x-Luciferase co-transfected HEK293 cells Inhibition - 7011-01_Antagonist_SinglePoint_HTS_Activity
|
|
Name: Fluorescence-based cell-based primary high throughput screening assay to identify pos...
Source: The Scripps Research Institute Molecular Screening Center
Target: muscarinic acetylcholine receptor M1 [Homo sapiens]
External Id: CHRM1_PAM_FLUO8_1536_1X%ACT PRUN
|
|
Name: Fluorescence polarization-based biochemical high throughput primary assay to identify...
Source: The Scripps Research Institute Molecular Screening Center
Target: RecName: Full=Sialate O-acetylesterase; AltName: Full=H-Lse; AltName: Full=Sialic acid-specific 9-O-acetylesterase; Flags: Precursor [Homo sapiens]
External Id: SIAE_INH_FP_1536_1X%INH PRUN
|
Total 130, Current Page 1 of 13
1
2
3
4
5
| HMS2229E04 |
| Benz(a)-1,3-benzodioxolo(4,5-g)quinolizinium,11,12-dihydro-8,9-dimethoxy |
| GNF-Pf-2355 |
| epiberberinium |
| 8,9-dimethoxy-11,12-dihydro-[1,3]dioxolo[4,5-h]isoquino[2,1-b]isoquinolinylium |
| 8,9-Dimethoxy-11,12-dihydro[1,3]dioxolo[4,5-h]isoquinolino[2,1-b]isoquinolin-13-ium |
| pseudo-Epiberberin |
| Epiberberin |

