Oxychloroaphine
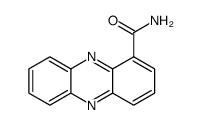
Oxychloroaphine structure
|
Common Name | Oxychloroaphine | ||
|---|---|---|---|---|
| CAS Number | 550-89-0 | Molecular Weight | 223.23000 | |
| Density | 1.371g/cm3 | Boiling Point | 526.1ºC at 760 mmHg | |
| Molecular Formula | C13H9N3O | Melting Point | 242ºC | |
| MSDS | N/A | Flash Point | 272ºC | |
Use of OxychloroaphineOxychloroaphine could be isolated from the bacterium Pantoea agglomerans naturally present in soil. Oxychloroaphine has broad-spectrum antifungal activity. Oxychloroaphine has cytotoxicity in a dose-dependent manner and induces apoptosis. Oxychloroaphine can be used in research of cancer[1][2]. |
| Name | phenazine-1-carboxamide |
|---|---|
| Synonym | More Synonyms |
| Description | Oxychloroaphine could be isolated from the bacterium Pantoea agglomerans naturally present in soil. Oxychloroaphine has broad-spectrum antifungal activity. Oxychloroaphine has cytotoxicity in a dose-dependent manner and induces apoptosis. Oxychloroaphine can be used in research of cancer[1][2]. |
|---|---|
| Related Catalog | |
| In Vitro | Oxychloroaphine (1-256 μM; 24 h) has cytotoxicity with IC50 values for A549, HeLa, and SW480 cancer cell lines between 32 and 40 μM[2]. Oxychloroaphine (1-150 μM; A549, HeLa, and SW480 cancer cell lines) causes cell membrane damage, leading to increase apoptosis and leakage of lactate dehydrogenase, and increases production of cytochrome c protein[2]. Oxychloroaphine (32 μM; A549 and SW480 cells) induces cycle arrest at G1 phase and induction of sub-G phase[2]. Oxychloroaphine (48 h; A549 cells) induces downregulation of antiapoptotic Bcl-2 protein and the activation of proapoptotic protein caspase-3 led to the cleavage of PARP[2]. Cell Viability Assay[2] Cell Line: A549, HeLa, and SW480 cancer cell lines Concentration: 1, 2, 4, 8, 16, 32, 64, 128, and 256 μM Incubation Time: 24 hours Result: Inhibited cell proliferative in a dose-dependent manner. |
| Density | 1.371g/cm3 |
|---|---|
| Boiling Point | 526.1ºC at 760 mmHg |
| Melting Point | 242ºC |
| Molecular Formula | C13H9N3O |
| Molecular Weight | 223.23000 |
| Flash Point | 272ºC |
| Exact Mass | 223.07500 |
| PSA | 68.87000 |
| LogP | 2.58220 |
| Index of Refraction | 1.76 |
| InChIKey | KPZYYKDXZKFBQU-UHFFFAOYSA-N |
| SMILES | NC(=O)c1cccc2nc3ccccc3nc12 |
| HS Code | 2933990090 |
|---|
|
~% 
Oxychloroaphine CAS#:550-89-0 |
| Literature: Lasseur;zit.bei Koegl;Postowsky Justus Liebigs Annalen der Chemie, 1930 , vol. 480, p. 280,289,291 |
|
~% 
Oxychloroaphine CAS#:550-89-0 |
| Literature: Borrero, Nicholas V.; Bai, Fang; Perez, Cristian; Duong, Benjamin Q.; Rocca, James R.; Jin, Shouguang; Huigens Iii, Robert W. Organic and Biomolecular Chemistry, 2014 , vol. 12, # 6 p. 881 - 886 |
|
~% 
Oxychloroaphine CAS#:550-89-0 |
| Literature: Borrero, Nicholas V.; Bai, Fang; Perez, Cristian; Duong, Benjamin Q.; Rocca, James R.; Jin, Shouguang; Huigens Iii, Robert W. Organic and Biomolecular Chemistry, 2014 , vol. 12, # 6 p. 881 - 886 |
|
~% 
Oxychloroaphine CAS#:550-89-0 |
| Literature: Toromanoff Annales de Chimie (Cachan, France), 1956 , vol. <13>1, p. 115,142 |
|
~% 
Oxychloroaphine CAS#:550-89-0 |
| Literature: Toromanoff Annales de Chimie (Cachan, France), 1956 , vol. <13>1, p. 115,142 |
|
~% 
Oxychloroaphine CAS#:550-89-0 |
| Literature: Dufraisse et al. Comptes Rendus Hebdomadaires des Seances de l'Academie des Sciences, 1952 , vol. 235, p. 920 Full Text Show Details Koegl; Postowsky Justus Liebigs Annalen der Chemie, 1930 , vol. 480, p. 280,286, 293 |
|
~% 
Oxychloroaphine CAS#:550-89-0 |
| Literature: Borrero, Nicholas V.; Bai, Fang; Perez, Cristian; Duong, Benjamin Q.; Rocca, James R.; Jin, Shouguang; Huigens Iii, Robert W. Organic and Biomolecular Chemistry, 2014 , vol. 12, # 6 p. 881 - 886 |
| HS Code | 2933990090 |
|---|---|
| Summary | 2933990090. heterocyclic compounds with nitrogen hetero-atom(s) only. VAT:17.0%. Tax rebate rate:13.0%. . MFN tariff:6.5%. General tariff:20.0% |
|
Name: GPR151 activator identification: cell-based high-throughput counter-screen assay
Source: The Scripps Research Institute Molecular Screening Center
Target: RecName: Full=Glucose-dependent insulinotropic receptor; AltName: Full=G-protein coupled receptor 119
External Id: GPR119_PHUNTER_AG_LUMI_1536_3X%ACT CSRUN1
|
|
Name: GPR151 activator identification: cell-based high-throughput confirmation assay
Source: The Scripps Research Institute Molecular Screening Center
Target: RecName: Full=G-protein coupled receptor 151; AltName: Full=G-protein coupled receptor PGR7; AltName: Full=GPCR-2037; AltName: Full=Galanin receptor 4; AltName: Full=Galanin-receptor-like protein; Short=GalRL
External Id: GPR151_PHUNTER_AG_LUMI_1536_3X%ACT CRUN1
|
|
Name: Antibacterial activity against Staphylococcus aureus after 2 days by disk assay
Source: ChEMBL
Target: Staphylococcus aureus
External Id: CHEMBL1028404
|
|
Name: Antibacterial activity against Bacillus cereus after 2 days by disk assay
Source: ChEMBL
Target: Bacillus cereus
External Id: CHEMBL1028402
|
|
Name: Antibacterial activity against methicillin-resistant Staphylococcus aureus after 16 t...
Source: ChEMBL
Target: Staphylococcus aureus
External Id: CHEMBL4041986
|
|
Name: Antibacterial activity against Micrococcus luteus after 2 days by disk assay
Source: ChEMBL
Target: Micrococcus luteus
External Id: CHEMBL1028403
|
|
Name: Cell-based high throughput primary assay to identify activators of GPR151
Source: The Scripps Research Institute Molecular Screening Center
Target: RecName: Full=G-protein coupled receptor 151; AltName: Full=G-protein coupled receptor PGR7; AltName: Full=GPCR-2037; AltName: Full=Galanin receptor 4; AltName: Full=Galanin-receptor-like protein; Short=GalRL
External Id: GPR151_PHUNTER_AG_LUMI_1536_1X%ACT
|
|
Name: AlphaScreen-based biochemical high throughput primary assay to identify activators of...
Source: The Scripps Research Institute Molecular Screening Center
Target: N/A
External Id: FBW7_ACT_ALPHA_1536_1X%ACT PRUN
|
|
Name: AlphaScreen-based biochemical high throughput primary assay to identify inhibitors of...
Source: The Scripps Research Institute Molecular Screening Center
External Id: MITF_INH_Alpha_1536_1X%INH PRUN
|
| α-Amidophenazin |
| phenazine-1-carboxylic acid amide |
| PCN |
| 1-Carbamoyl-phenazin |
| Phenazine-1-carboxamide |
| Phenazin-1-carbamid |


 CAS#:2876-22-4
CAS#:2876-22-4