H-Met-Ala-Ser-OH
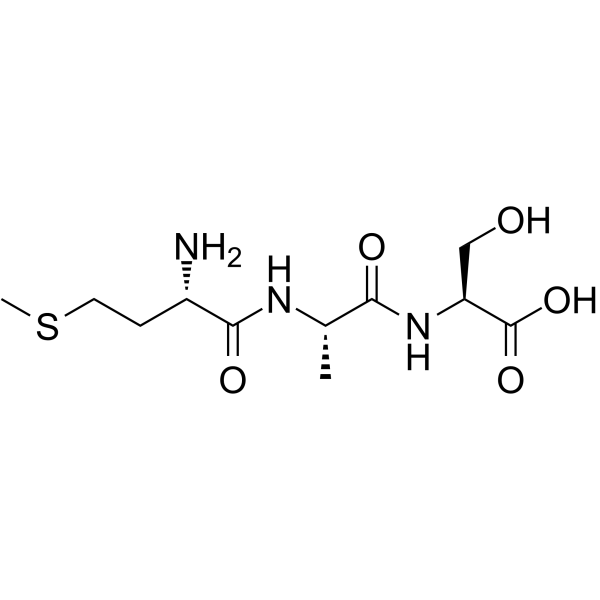
H-Met-Ala-Ser-OH structure
|
Common Name | H-Met-Ala-Ser-OH | ||
|---|---|---|---|---|
| CAS Number | 17351-33-6 | Molecular Weight | 307.36700 | |
| Density | N/A | Boiling Point | N/A | |
| Molecular Formula | C11H21N3O5S | Melting Point | N/A | |
| MSDS | Chinese USA | Flash Point | N/A | |
| Symbol |

GHS07 |
Signal Word | Warning | |
|
Interactions of platinum(II) complexes with sulfur-containing peptides studied by electrospray ionization mass spectrometry and tandem mass spectrometry.
Rapid Commun. Mass Spectrom. 19 , 1031-1040, (2005) Reactions of two platinum(II) complexes, cis-[Pt(NH3)2(H2O)2]2+ (Pt1) and cis-[Pt(en)(H2O)2]2+ (Pt2), with several sulfur-containing peptides, have been investigated by electrospray ionization mass spectrometry (ESI-MS) and tandem mass spectrometry (MS/MS). T... |
|
|
Molecular cloning and characterization of a new peptide deformylase from human pathogenic bacterium Helicobacter pylori.
Biochem. Biophys. Res. Commun. 319 , 1292-1298, (2004) Helicobacter pylori is a gram-negative pathogenic bacterium, which is associated with peptic ulcer disease and gastric cancer. It is urgent to discover novel drug targets for appropriate antimicrobial agents against this human pathogen. In bacteria, peptide d... |
|
|
Crystal structure of an EfPDF complex with Met-Ala-Ser based on crystallographic packing.
Biochem. Biophys. Res. Commun. 381 , 630-633, (2009) PDF (peptide deformylase) plays a critical role in the production of mature proteins by removing the N-formyl polypeptide of nascent proteins in the prokaryote cell system. This protein is essential for bacterial growth, making it an attractive target for the... |
|
|
Iron center, substrate recognition and mechanism of peptide deformylase.
Nat. Struct. Biol. 5 , 1053-1058, (1998) Eubacterial proteins are synthesized with a formyl group at the N-terminus which is hydrolytically removed from the nascent chain by the mononuclear iron enzyme peptide deformylase. Catalytic efficiency strongly depends on the identity of the bound metal. We ... |