5-Hydroxymethyluracil

5-Hydroxymethyluracil structure
|
Common Name | 5-Hydroxymethyluracil | ||
|---|---|---|---|---|
| CAS Number | 4433-40-3 | Molecular Weight | 142.113 | |
| Density | 1.4±0.1 g/cm3 | Boiling Point | 530.4ºC at 760mmHg | |
| Molecular Formula | C5H6N2O3 | Melting Point | >300 °C(lit.) | |
| MSDS | Chinese USA | Flash Point | 274.6ºC | |
Use of 5-Hydroxymethyluracil5-Hydroxymethyluracil is a product of oxidative DNA damage. 5-Hydroxymethyluracil can be used as a potential epigenetic mark enhancing or inhibiting transcription with bacterial RNA polymerase. |
| Name | 5-hydroxymethyluracil |
|---|---|
| Synonym | More Synonyms |
| Description | 5-Hydroxymethyluracil is a product of oxidative DNA damage. 5-Hydroxymethyluracil can be used as a potential epigenetic mark enhancing or inhibiting transcription with bacterial RNA polymerase. |
|---|---|
| Related Catalog | |
| Target |
Human Endogenous Metabolite |
| References |
| Density | 1.4±0.1 g/cm3 |
|---|---|
| Boiling Point | 530.4ºC at 760mmHg |
| Melting Point | >300 °C(lit.) |
| Molecular Formula | C5H6N2O3 |
| Molecular Weight | 142.113 |
| Flash Point | 274.6ºC |
| Exact Mass | 142.037842 |
| PSA | 86.47000 |
| LogP | -0.50 |
| Index of Refraction | 1.528 |
| InChIKey | JDBGXEHEIRGOBU-UHFFFAOYSA-N |
| SMILES | O=c1[nH]cc(CO)c(=O)[nH]1 |
| Storage condition | Refrigerator |
CHEMICAL IDENTIFICATION
HEALTH HAZARD DATAACUTE TOXICITY DATAMUTATION DATA
|
| Personal Protective Equipment | Eyeshields;Gloves;type N95 (US);type P1 (EN143) respirator filter |
|---|---|
| Hazard Codes | Xi: Irritant; |
| Safety Phrases | S22-S24/25 |
| RIDADR | NONH for all modes of transport |
| WGK Germany | 3 |
| RTECS | YR0513000 |
| HS Code | 2933599090 |
| Precursor 10 | |
|---|---|
| DownStream 10 | |
| HS Code | 2933599090 |
|---|---|
| Summary | 2933599090. other compounds containing a pyrimidine ring (whether or not hydrogenated) or piperazine ring in the structure. VAT:17.0%. Tax rebate rate:13.0%. . MFN tariff:6.5%. General tariff:20.0% |
|
Measurement of urinary excretion of 5-hydroxymethyluracil in human by GC/NICI/MS: correlation with cigarette smoking, urinary TBARS and etheno DNA adduct.
Toxicol. Lett. 155(3) , 403-10, (2005) 5-Hydroxymethyluracil (5-HMU) is derived from radiation in addition to endogenous oxidative DNA damage and it is one of the most abundant DNA adducts. Human 5-HMU-DNA-glycosylase has been shown to rep... |
|
|
Mutational analysis of the damage-recognition and catalytic mechanism of human SMUG1 DNA glycosylase.
Nucleic Acids Res. 32(17) , 5291-302, (2004) Single-strand selective monofunctional uracil-DNA glycosylase (SMUG1), previously thought to be a backup enzyme for uracil-DNA glycosylase, has recently been shown to excise 5-hydroxyuracil (hoU), 5-h... |
|
|
Qualitative and quantitative analyses of the decomposition products that arise from the exposure of thymine to monochromatic ultrasoft X rays and 60Co gamma rays in the solid state.
Radiat. Res. 161(4) , 442-50, (2004) HPLC analyses of condensed thymine irradiated with monochromatic synchrotron ultrasoft X rays in the energy region around nitrogen and oxygen K-shell edges were performed. Cobalt-60 gamma rays were us... |
|
Name: Discovery of Small Molecules to Inhibit Human Cytomegalovirus Nuclear Egress
Source: ICCB-Longwood/NSRB Screening Facility, Harvard Medical School
Target: HCMV UL50
External Id: HMS1262
|
|
Name: Chemical Probes of Kaposi's Sarcoma Herpes Virus Latent Infection
Source: ICCB-Longwood/NSRB Screening Facility, Harvard Medical School
Target: ORF 73 [Human herpesvirus 8 type M]
External Id: HMS791
|
|
Name: Competitive inhibition of human erythrocytes CA1 using 4-nitrophenylacetate as substr...
Source: ChEMBL
Target: Carbonic anhydrase 1
External Id: CHEMBL3602571
|
|
Name: Inhibition of human erythrocytes CA1 using 4-nitrophenylacetate as substrate by ester...
Source: ChEMBL
Target: Carbonic anhydrase 1
External Id: CHEMBL3602569
|
|
Name: A screen for compounds that inhibit the activity of LtaS in Staphylococcus aureus
Source: ICCB-Longwood/NSRB Screening Facility, Harvard Medical School
External Id: HMS979
|
|
Name: Inhibition of human erythrocytes CA2 using 4-nitrophenylacetate as substrate by ester...
Source: ChEMBL
Target: Carbonic anhydrase 2
External Id: CHEMBL3602570
|
|
Name: Cell-based high throughput primary assay to identify activators of GPR151
Source: The Scripps Research Institute Molecular Screening Center
Target: RecName: Full=G-protein coupled receptor 151; AltName: Full=G-protein coupled receptor PGR7; AltName: Full=GPCR-2037; AltName: Full=Galanin receptor 4; AltName: Full=Galanin-receptor-like protein; Short=GalRL
External Id: GPR151_PHUNTER_AG_LUMI_1536_1X%ACT
|
|
Name: Stabilization of p53 in human papillomavirus-positive cells
Source: ICCB-Longwood/NSRB Screening Facility, Harvard Medical School
Target: Cellular tumor antigen p53
External Id: HMS1485
|
|
Name: A screen for compounds that inhibit viral RNA polymerase binding and polymerization a...
Source: ICCB-Longwood/NSRB Screening Facility, Harvard Medical School
Target: Chain A, Poliovirus Polymerase With Gtp
External Id: HMS750
|
|
Name: Primary qHTS for inhibitors of NSP2Pro chikungunya virus (CHIKV)
Source: NCGC
External Id: APP-Toga-CHIKV-nsp2-p
|
| 5-(hydroxymethyl)-1H-pyrimidine-2,4-dione |
| EINECS 224-636-5 |
| 5-(hydroxymethyl)pyrimidine-2,4(1H,3H)-dione |
| 5-hydroxymethyluracil |
| MFCD00056024 |
| 5-(hydroxymethyl)-1,3-dihydropyrimidine-2,4-dione |
| 5-(Hydroxymethyl)uracil |
| 5-(Hydroxymethyl)-2,4(1H,3H)-pyrimidinedione |
 CAS#:50-00-0
CAS#:50-00-0 CAS#:66-22-8
CAS#:66-22-8 CAS#:65-71-4
CAS#:65-71-4 CAS#:89179-86-2
CAS#:89179-86-2 CAS#:3590-48-5
CAS#:3590-48-5 CAS#:107-21-1
CAS#:107-21-1 CAS#:825649-02-3
CAS#:825649-02-3 CAS#:876495-24-8
CAS#:876495-24-8 CAS#:856152-76-6
CAS#:856152-76-6![5-((benzo[d]oxazol-2-ylthio)methyl)pyrimidine-2,4(1H,3H)-dione Structure](https://image.chemsrc.com/caspic/055/84345-71-1.png) CAS#:84345-71-1
CAS#:84345-71-1![[2,4-bis(trimethylsilyloxy)pyrimidin-5-yl]methanol structure](https://image.chemsrc.com/caspic/385/111878-20-7.png) CAS#:111878-20-7
CAS#:111878-20-7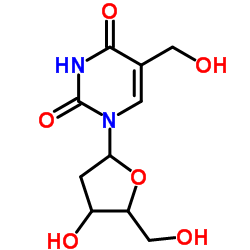 CAS#:5116-24-5
CAS#:5116-24-5 CAS#:7627-38-5
CAS#:7627-38-5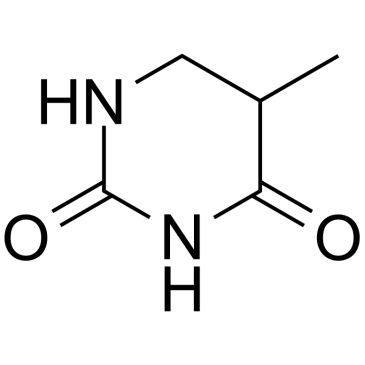 CAS#:696-04-8
CAS#:696-04-8 CAS#:108141-35-1
CAS#:108141-35-1 CAS#:1195-08-0
CAS#:1195-08-0![2,4(1H,3H)-Pyrimidinedione,5-[(4-hydroxyphenyl)methyl] structure](https://image.chemsrc.com/caspic/316/17187-50-7.png) CAS#:17187-50-7
CAS#:17187-50-7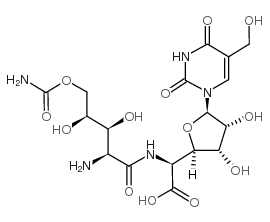 CAS#:19396-06-6
CAS#:19396-06-6
